Mouse Mesangial Cells
Cat.No.: CSC-C9368W
Species: Mouse
Source: Kidney
Morphology: Polygonal
Cell Type: Mesangial Cell
- Specification
- Background
- Scientific Data
- Q & A
- Customer Review
Mouse Mesangial Cells (MMCs) are a population of perivascular cells that reside within the central stalk of the kidney glomerulus. Permeating the central stalk are capillaries responsible for filtration of blood in order to create urine. Mesangial cells are the predominant cell-type in this structure referred to as glomerular mesangium, and reside within an extracellular matrix that they synthesize and modify themselves. Morphologically, MMCs have a "stellate" or spindle-shape appearance. They exhibit contractile ability and are able to contract and expand the surface area of glomerular capillaries by responding to circulating vasoactive factors such as Angiotensin II and Endothelin-1. Due to this reason, mesangial cells play a crucial role in setting Glomerular Filtration Rate (GFR).
Mesangial cells also function as immune and metabolic sensors of the kidney. They are capable of phagocytosing macromolecules and immune complexes that pass into the mesangium and prevent them from clogging the filtration system. During disease states, MMCs often switch from a quiescent phenotype to an activated proliferative phenotype. Conditions that can involve mesangial cell activation include Diabetic Nephropathy, Glomerulonephritis, and many other diseases affecting the kidney. Activated mesangial cells often overproduce inflammatory cytokines (IL-6, TNF-α) and profibrotic growth factors such as TGF-β. This overproduction causes mesangial expansion and excess production of extracellular matrix which leads to glomerulosclerosis and eventually chronic kidney disease (CKD).
Determination of Reference Genes as a Quantitative Standard for Gene Expression Analysis in Mouse Mesangial Cells Stimulated with TGF-Β
Reverse transcription-quantitative polymerase chain reaction (RT-PCR) is a standard technique for gene expression analysis, but selecting appropriate reference genes (housekeeping genes, HKG) is challenging. dos Santos Bronel et al. identified the best HKG for an in vitro model using mouse mesangial cells (MMCs) stimulated with TGF-β.
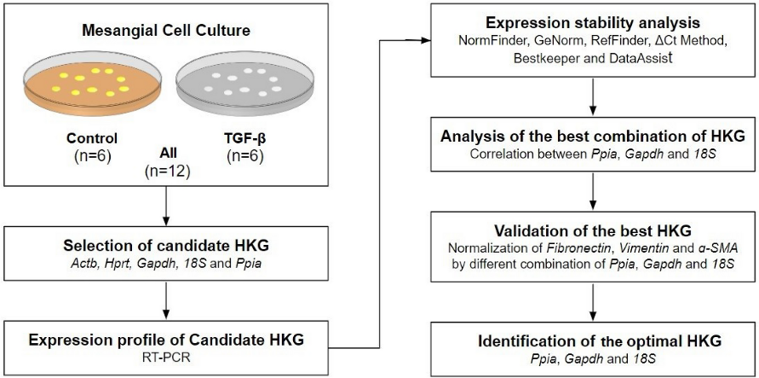
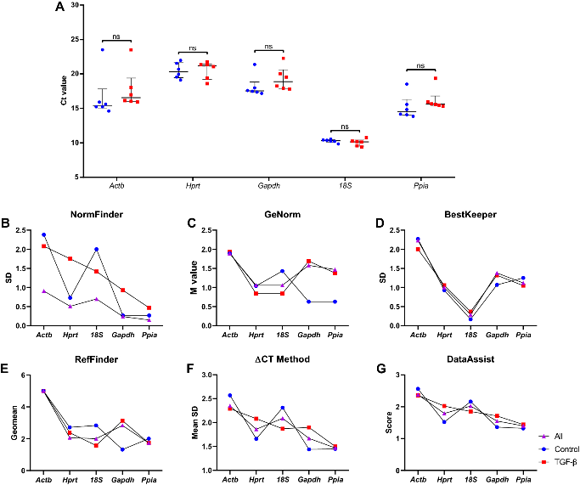
To identify the best housekeeping gene (HKG) by RT-PCR, they used a stepwise strategy. Figure 1 summarizes the workflow. Samples were divided into three sets: control cells (n = 6), TGF-β-treated cells (n = 6), and All (n = 12). Ct values of the five candidate HKGs ranged from 23.511 to 9.387, shown as median (interquartile range). Ct values were inversely correlated with gene expression levels. Hprt showed the highest Ct value [20.876 (2.05)], indicating that it was the least expressed among the five candidate HKGs. In contrast, 18S had the lowest Ct value [10.232 (0.50)], suggesting that it was the most abundantly expressed HKG. Gapdh [17.948 (2.41)], Actb [15.986 (2.44)], and Ppia [15.514 (1.54)] were moderately expressed. The median Ct value for each HKG is shown in Fig. 2A. No significant difference was observed between control cells and TGF-β-treated cells.
Ask a Question
Write your own review