The 8 Costliest Mistakes in Preclinical CYP Phenotyping
You run liver microsome assays. Everything looks fine. But six months later, your PK study fails-unexpected drug interactions, nonlinear clearance, or a competitor beats you to IND. What went wrong?
In discovery and preclinical development, CYP phenotyping isn't a checkbox-it's a make-or-break step. Avoid these eight mistakes, and you deliver data that drives programs forward instead of delaying them.
Quick Reference: The 8 Pitfalls
| Mistake | Where It Happens | Potential Impact |
|---|---|---|
| 1. Wrong test system | Early screening (microsomes vs. hepatocytes) | Missed clearance pathways, false negatives in DDI |
| 2. Ignoring enzyme kinetics | IC50 determinations | Misranked DDI risk, inaccurate clinical predictions |
| 3. Single-point IC50s | High-throughput screening | 10-100× error in potency; false negatives |
| 4. Overlooking transporters | Hepatocyte uptake/metabolism studies | Misestimated clearance, false DDI risk |
| 5. Static DDI predictions | IND packages | Missed mechanism-based inhibition/induction; delayed FDA approval |
| 6. Mismatched animal models | In vivo PK species selection | Unpredictable human PK; wasted GLP tox |
| 7. Poor probe selection | Cocktail inhibition studies | Non-interpretable data; repeated costly studies |
| 8. Data silos | Separate ADME, PK, efficacy reports | Missed clinical risk, lost business |
Mistake 1: Using the Wrong Test System
The error: Running all CYP studies in microsomes because they are cheap and fast.
Reality: Microsomes lack cellular machinery-they miss transporter effects, enzyme induction, and some forms of TDI. For example, midazolam clearance was underpredicted 5-fold when microsomes missed CYP3A4/5 contribution in hepatocytes.
Fix: Use a tiered system:
| Question | System | Why |
|---|---|---|
| Metabolic stability & CYP-specific phenotyping | Liver microsomes + recombinant CYPs | Cost-effective, CYP-specific |
| Induction potential (mRNA/activity) | Primary human hepatocytes | Requires nuclear receptors (PXR, CAR, AhR) |
| Mechanism-based inhibition / TDI | Fresh hepatocytes or microsomes with preincubation | Captures mechanism-based inactivation |
| Integrated clearance (uptake + metabolism) | Suspended/plated hepatocytes | Accounts for transporter-enzyme interplay |
Mistake 2: Ignoring Enzyme Kinetics
The error: Reporting IC50 values without substrate correction or inhibition mechanism.
Reality: IC50 changes with substrate concentration relative to Km. For competitive inhibitors:
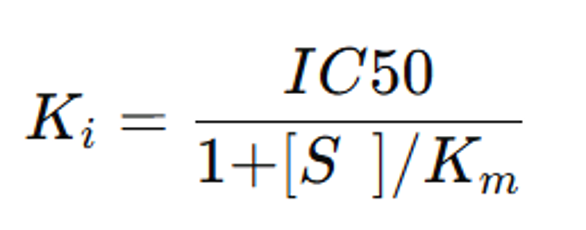
Note: Formula valid for reversible, competitive inhibitors only. Non-competitive or TDI requires separate analysis.
Fix:
- Run 8-point inhibition curves in duplicate
- Correct for plasma protein binding (fu)
- Calculate [I]/Ki ratios: >0.1 = weak DDI risk; >1 = strong risk
- Flag non-competitive kinetics for follow-up
Mistake 3: Relying on Single-Point IC50s
The error: Screening at a single inhibitor concentration.
Reality: Partial inhibition, steep/shallow curves, or low-concentration activation can be missed. Compounds labeled "inactive at 10 μM" may have potent IC50s at 0.1 μM.
Fix:
- Use 8-point half-log curves with positive controls (ketoconazole, quinidine)
- Report Hill slope; flag >2 or <0.5
- For HTS, use 3-point screens only for initial hits; confirm with full curves
Mistake 4: Overlooking Transporters
The error: Treating hepatocytes as "CYP in a dish."
Reality: Hepatic uptake often limits clearance (OATP1B1/1B3, NTCP). Ignoring uptake vs. metabolism confuses clearance prediction and DDI risk.
Fix:
- Separate uptake assays (37°C vs 4°C)
- Use transporter inhibitors to isolate metabolic contribution
- For CYP phenotyping, ensure substrate concentration exceeds uptake Km
- Flag high-clearance compounds for transporter contribution assessment
Mistake 5: Static DDI Predictions Only
The error: Submitting only reversible inhibition data for IND.
Reality: FDA guidance (2020) requires reversible inhibition, TDI, and induction. Static [I]/Ki misses 30-50% of clinical DDIs (e.g., clarithromycin-CYP3A4, rifampin induction).
Fix:
- TDI assay: Preincubate with NADPH, measure IC50 shift ≥1.5×
- Induction assay: Treat hepatocytes 48-72 hr, measure mRNA/activity; use positive controls (rifampin, omeprazole, phenobarbital)
- IVIVE: Estimate R-value, kdeg, Emix; flag for clinical study design
Note: PBPK optional-static models are sufficient for IND if applied correctly
Mistake 6: Mismatched Animal Models
The error: Defaulting to rat PK studies without checking CYP orthology.
Reality: Rats differ from humans in CYP isoforms and induction. For instance, a CYP2D6 substrate in humans may clear via CYP3A in dogs.
| Human CYP | Preferred Models | Avoid |
|---|---|---|
| CYP3A4 | Dog, monkey, humanized mice | Rat |
| CYP2D6 | Monkey, humanized mice | Dog (non-functional) |
| CYP2C9 | Monkey | Rat |
| CYP1A2 | Most species (consider induction differences) | - |
Cross-species in vitro comparison before GLP tox ensures PK/PD relevance.
Mistake 7: Poor Probe Selection
The error: Using non-selective probes or poorly separated metabolites.
Reality: Testosterone (CYP3A4) may also hit CYP2C → uninterpretable data. Studies often repeated at $50K+ cost.
Fix: Use validated, selective probes:
| CYP | Probe | Metabolite | LC-MS/MS Notes |
|---|---|---|---|
| CYP1A2 | Phenacetin | Acetaminophen | Watch phase II |
| CYP2B6 | Bupropion | Hydroxybupropion | Chiral separation if needed |
| CYP2C8 | Amodiaquine | Desethylamodiaquine | Avoid MPPG interference |
| CYP2C9 | Diclofenac | 4'-OH-diclofenac | Stable at low pH |
| CYP2C19 | Omeprazole | 5-OH-omeprazole | Sulfide interference |
| CYP2D6 | Dextromethorphan | Dextrorphan | Separate from levorphan |
| CYP3A4/5 | Midazolam | 1'-OH-midazolam | Prefer 1' metabolite |
Validate selectivity with isoform-specific inhibitors (furafylline, quinidine, ketoconazole).
Mistake 8: Siloed Data Delivery
The error: Providing ADME, PK, and efficacy as separate reports.
Reality: Clients often miss critical risk combinations: high plasma protein binding + low solubility + CYP3A4 TDI = high clinical DDI risk.
Fix:
- Provide integrated risk assessment, flag showstoppers early
- Example: "CLint > 80% hepatic blood flow, fu < 0.01, CYP3A4 TDI positive → high clinical DDI risk. Recommend CYP3A4 phenotyping + transporter assessment + dedicated DDI package."
- Offer follow-up consultation to interpret results in program context
Business angle: Proactive risk assessment builds trust and repeat business.
CYP phenotyping isn't about perfect prediction-it's about informed risk management. Avoid these eight mistakes, and your data drives programs forward instead of delaying them.
Creative Bioarray Relevant Recommendations
| Products & Services | Description |
|---|---|
| CYP and UGT Reaction Phenotyping Assay | Creative Bioarray helps provide CYP and UGT reaction phenotyping assay, with years of experience and excellent scientific team to provide you with quality service. |
| Drug-Drug Interaction | Creative Bioarray provides a range of high-quality drug-drug interaction services to meet FDA and EMA guidance. |