A Complete Guide to CYP Reaction Phenotyping in 2026
CYP reaction phenotyping is a fundamental concept in drug metabolism and pharmacokinetics (DMPK), yet it is often misunderstood or oversimplified. As drug candidates progress from early discovery into preclinical development, understanding which cytochrome P450 (CYP) enzymes are responsible for their metabolism becomes critical for predicting clearance, variability, and drug-drug interaction (DDI) risk.
In simple terms, CYP reaction phenotyping answers one key question:
Which CYP enzymes metabolize a given drug, and to what extent?
This information plays a central role in lead optimization, regulatory strategy, and DDI risk assessment. In fact, poorly designed or misinterpreted reaction phenotyping studies are a common reason why DDI strategies need to be revisited later-often under regulatory pressure.
Why Is CYP Reaction Phenotyping Important?
CYP reaction phenotyping is important because it defines which metabolic risks truly matter during drug development. By identifying the CYP enzymes responsible for a drug's clearance, reaction phenotyping helps:
- Determine whether a compound is vulnerable to clinically relevant drug-drug interactions
- Prioritize CYP inhibition or induction studies based on actual metabolic contribution
- Support rational regulatory and clinical DDI strategies
Without reaction phenotyping, DDI assessments often rely on assumptions rather than mechanistic evidence, increasing the risk of late-stage surprises or regulatory questions.
How Is CYP Reaction Phenotyping Performed?
CYP reaction phenotyping is not a single assay, but a systematic experimental strategy designed to identify and quantify the contribution of individual CYP enzymes to drug metabolism. Regulatory agencies and industry best practices consistently recommend using multiple complementary in vitro approaches to ensure reliable interpretation.
In practice, reaction phenotyping integrates data from human-derived systems, selective perturbation methods, and quantitative scaling to establish a mechanistic basis for downstream DDI risk assessment.
Human-Derived In Vitro Systems as the Experimental Foundation
Most CYP reaction phenotyping studies are conducted in human liver microsomes (HLM) or human hepatocytes, as these systems best reflect clinically relevant CYP expression profiles.
- Human liver microsomes (HLM) provide a concentrated source of CYP enzymes and are widely used for initial phenotyping due to their reproducibility and experimental simplicity.
- Human hepatocytes retain additional cellular context, including uptake transporters and phase II enzymes, and may be used when metabolite formation or intracellular exposure is a concern.
Selection of the biological system influences both sensitivity and interpretability and should be aligned with the intended application of the data.
1. Chemical Inhibition Approach
The chemical inhibition method is one of the most established approaches for CYP reaction phenotyping and is explicitly discussed in regulatory guidance documents.
Selective chemical inhibitors are used to suppress individual CYP enzymes in HLM or hepatocytes. A reduction in intrinsic clearance or metabolite formation following inhibition indicates the involvement of the targeted CYP enzyme.
Typical workflow
- Incubate the test compound under linear metabolic conditions
- Apply validated concentrations of isoform-selective CYP inhibitors
- Compare metabolic rates with and without inhibitors
- Estimate relative enzyme contribution based on percent inhibition
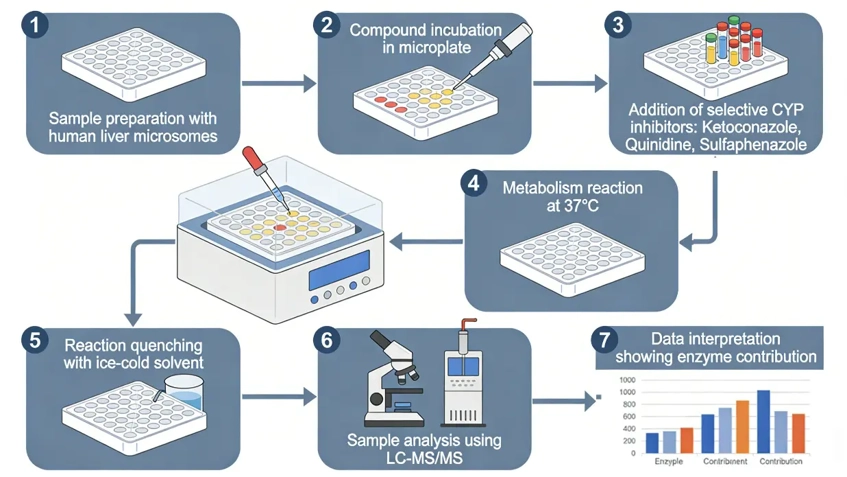
Critical considerations
- Inhibitor concentrations must balance selectivity and completeness
- Many inhibitors exhibit partial cross-reactivity at higher concentrations
- Time-dependent inhibition of the enzyme by the test compound itself must be ruled out
Because of these limitations, chemical inhibition is best suited for qualitative or semi-quantitative enzyme attribution rather than definitive fm estimation.
2. Recombinant CYP Enzyme Approach
Recombinant CYP phenotyping uses individually expressed human CYP isoforms to directly assess metabolic capability.
The compound is incubated with recombinant CYP enzymes under standardized conditions, allowing direct measurement of intrinsic clearance or metabolite formation by each isoform.
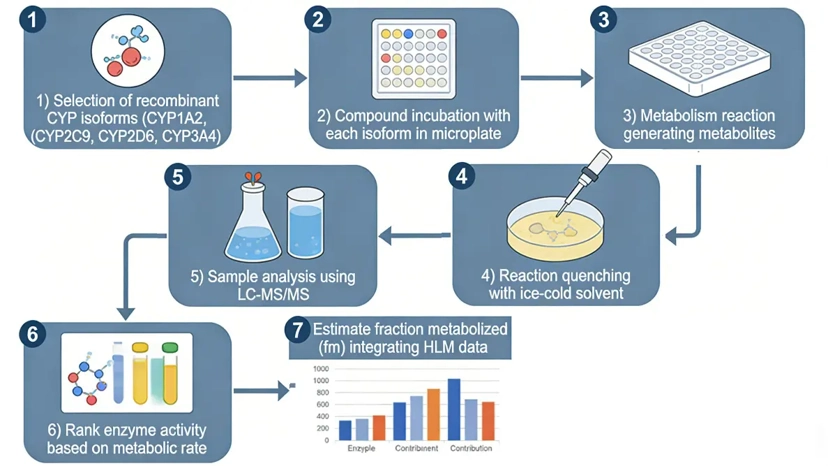
Applications
- Identification of metabolically competent CYP enzymes
- Ranking of CYP isoforms based on catalytic efficiency
- Detection of minor metabolic pathways not apparent in HLM
Limitations
- Recombinant systems do not reflect native hepatic enzyme abundance
- Activity may be exaggerated for low-expression CYP isoforms
- Data require scaling to be physiologically meaningful
As highlighted in multiple industry reviews, recombinant CYP data should be interpreted as enzyme capability, not enzyme contribution.
3. Quantitative Scaling and fm Estimation
To translate in vitro findings into meaningful DDI risk assessments, reaction phenotyping studies aim to estimate the fraction metabolized (fm) by each CYP enzyme.
fm values are typically derived by integrating:
- Chemical inhibition data from HLM or hepatocytes
- Recombinant CYP activity normalized by relative enzyme abundance
- Total intrinsic clearance measurements
Several scaling approaches are described in the literature, including:
- Relative activity factor (RAF) methods
- Inter-system extrapolation factor (ISEF) approaches
Regardless of the method used, regulatory agencies expect fm estimates to be mechanistically justified and internally consistent.
4. Use of Multiple Complementary Approaches
Current best practices emphasize that no single method is sufficient for robust CYP reaction phenotyping.
Instead, a convergent strategy is recommended:
- Chemical inhibition to assess enzyme contribution in a physiologically relevant matrix
- Recombinant CYP assays to confirm metabolic capability
- Quantitative integration to estimate fm values
This integrated approach reduces uncertainty and improves confidence in downstream DDI risk categorization.
Alignment with regulatory and industry guidance
Both FDA and EMA guidance documents emphasize the importance of understanding metabolic pathways early in development. While specific experimental designs are not mandated, reaction phenotyping data are expected to:
- Support selection of CYP enzymes for inhibition and induction studies
- Inform clinical DDI study design
- Provide mechanistic context for regulatory submissions
Well-designed reaction phenotyping studies therefore function as a foundational dataset, rather than a stand-alone experiment.
5. Common Experimental Pitfalls Highlighted in the Literature
Published analyses consistently identify several recurring issues:
- Overreliance on a single inhibitor or method
- Misinterpretation of partial inhibition data
- Inconsistent incubation conditions across assays
- Inadequate consideration of time-dependent inhibition
Addressing these challenges requires both technical expertise and experience with regulatory expectations, which is why reaction phenotyping is frequently outsourced to specialized DMPK laboratories.
Creative Bioarray Relevant Recommendations
| Products & Services | Description |
|---|---|
| CYP and UGT Reaction Phenotyping Assay | Creative Bioarray helps provide CYP and UGT reaction phenotyping assay, with years of experience and excellent scientific team to provide you with quality service. |
| Drug-Drug Interaction | Creative Bioarray provides a range of high-quality drug-drug interaction services to meet FDA and EMA guidance. |