DNA Laddering Assay
One of the distinctive biochemical features of apoptosis is the cleavage of DNA by a specific nuclease called caspase-activated DNase (CAD). During apoptosis, the DNA strand is cleaved by CAD and generates a number of DNA fragmentations of 180-200 base pairs known as DNA ladders. These DNA fragments can be extracted from the cells and visualized using agarose gel electrophoresis. Agarose gel electrophoresis is a simple technique but generally able to provide robust results. In order to compare the degree of apoptosis between experimental samples, more quantitative methods need to be combined.
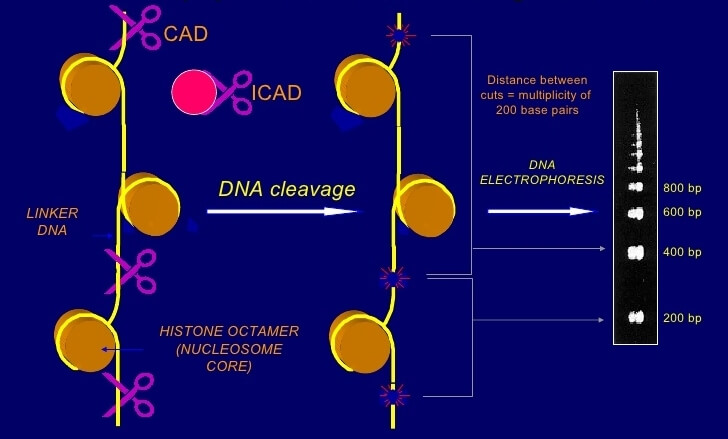 Figure 1. Apoptosis-induce DNA fragmentation.
Figure 1. Apoptosis-induce DNA fragmentation.
Materials
- TAE
- Agarose
- Gel loading buffer
- DNA size markers
- Ethidium bromide
- Proteinase K
- RNase cocktail
- TES lysis buffer
Equipment
- Standard gel electrophoresis equipment
- Refrigerated cell centrifuge
- UV transilluminator
- DC power supply
- Heating block
- Microfuge
Assay Procedure
- Prepare 0.5 mL of cell suspension (The number of cells is no less than 5 x 105 and no more than 5 x 106).
Note: No less than 5 x 105, otherwise DNA will not be detected, and no more than 5 x 106, to avoid difficult handling of insoluble DNA. - Centrifuge cells at 2000 rpm at 4°C for 10 min.
- Add 0.5 mL of TES lysis buffer and vortex vigorously.
Note: This procedure allows the release of fragmented chromatin from nuclei, after cell lysis and disruption of the nuclear structure. - Add 20 µL of RNase cocktail and mix well by flipping the tip of the tube.
- Incubate for 30-120 minutes at 37°C.
- Add 20 µL of proteinase K and mix by flipping the tip of the tube.
- Incubate at 50°C for at least 90 minutes or, if preferred, overnight.
- Mix the samples of DNA with loading buffer.
Note: The addition of loading buffer to samples allows to load gel wells more easily and to monitor the run of samples. - Load 10-20 µL of DNA samples to each well of a standard 1% agarose gel containing 0.5 µg/mL ethidium bromide.
Note: Ethidium bromide is a potential carcinogen, please wear gloves and handle with care.
Instead of directly adding ethidium bromide into the agarose gel, DNA can also be visualized by staining the gel after electrophoresis in 1 µg/mL ethidium bromide (containing TAE buffer) for 1 hour. - Run the gel at a low voltage, which improves resolution of DNA fragments (35 V for 4 hours or until loading dye has run two-thirds of the way down the gel).
Note: During electrophoresis, it is possible to monitor the migration of samples by following the migration of bromophenol blue dye contained in the loading buffer. - Visualize the DNA by a UV light source and documented by photography.
Note: Wear eye and skin protection when UV are on.
Apoptotic cells can form a distinct DNA ladder, while necrotic samples may generate a smear pattern (or no DNA splitting). DNA from viable cells will remain at the top of the gel as a high-molecular-weight band.
References
- Bortner, C.D. et al.; The role of DNA fragmentation in apoptosis. Trends Cell Biol. 1995, 5: 21.
- Duke, R.C. et al.; Endogenous endonuclease-induced DNA fragmentation: an early event in cell-mediated cytolysis. Proc. Natl. Acad. Sci. U.S.A. 1986, 80: 6361.
Related Sections
Cell Services:
Cell Line Testing and Assays:
For research use only. Not for any other purpose.
Resources
- FAQ
- Protocol
- Cell Culture Guide
- Technical Bulletins
-
Explore & Learn
-
Cell Biology
- Monocytes vs. Macrophages
- How to Detect and Remove Endotoxins in Biologics?
- Comparison of Different Methods to Measure Cell Viability
- What Are Myeloid Cell Markers?
- Contamination of Cell Cultures & Treatment
- Generation and Applications of Neural Stem Cells
- Stem Cell Markers
- Quantification of Cytokines
- T Cell Activation and Expansion
- Comparison of the MSCs from Different Sources
- How to Isolate and Analyze Tumor-Infiltrating Leukocytes?
- Guidelines for Cell Banking to Ensure the Safety of Biologics
- Cell Cryopreservation Techniques and Practices
- CFU Assay for Hematopoietic Cell
- Techniques for Cell Separation
- Circulating Tumor Cells as Cancer Biomarkers in the Clinic
- Biomarkers and Signaling Pathways in Tumor Stem Cells
- How to Start Your Culture: Thawing Frozen Cells
- Critical Quality Attributes and Assays for Induced Pluripotent Stem Cells
- What Is Cell Proliferation and How to Analyze It?
- Direct vs. Indirect Cell-Based ELISA
- STR Profiling—The ID Card of Cell Line
- Comparison of Several Techniques for the Detection of Apoptotic Cells
- How to Assess the Migratory and Invasive Capacity of Cells?
- What Cell Lines Are Commonly Used in Biopharmaceutical Production?
- Mesenchymal Stem Cells: A Comprehensive Exploration
- What are the Differences Between M1 and M2 Macrophages?
- Cryopreservation of Cells Step by Step
- What are PBMCs?
- Enrichment, Isolation and Characterization of Circulating Tumor Cells (CTCs)
- Neural Differentiation from Induced Pluripotent Stem Cells
- Isolation, Expansion, and Analysis of Natural Killer Cells
- Tumor Stem Cells: Identification, Isolation and Therapeutic Interventions
- Multi-Differentiation of Peripheral Blood Mononuclear Cells
- Tips For Cell Cryopreservation
- How to Decide Between 2D and 3D Cell Cultures?
- Organoid Differentiation from Induced Pluripotent Stem Cells
- Human Primary Cells: Definition, Assay, Applications
- What are Mesothelial Cells?
- How to Scale Up Single-Cell Clones?
- T Cell, NK Cell Differentiation from Induced Pluripotent Stem Cells
- Major Problems Caused by the Use of Uncharacterized Cell Lines
- IL-12 Family Cytokines and Their Immune Functions
- Cell Culture Medium
- How to Eliminate Mycoplasma From Cell Cultures?
- CHO Cell Line Development
- Troubleshooting Cell Culture Contamination: A Comprehensive Guide
- Strategies for Enrichment of Circulating Tumor Cells (CTCs)
- Unveiling the Molecular Secrets of Adipogenesis in MSCs
- How to Handle Mycoplasma in Cell Culture?
- How to Isolate PBMCs from Whole Blood?
- Spheroid vs. Organoid: Choosing the Right 3D Model for Your Research
- From Collection to Cure: How ACT Works in Cancer Immunotherapy
- Cell-Based High-Throughput Screening Techniques
- From Blur to Clarity: Solving Resolution Limits in Live Cell Imaging
- Overview of Cell Apoptosis Assays
- Live Cell Imaging: Unveiling the Dynamic World of Cellular Processes
- Optimization Strategies of Cell-Based Assays
- Live Cell Imaging: Unveiling the Dynamic World of Cellular Processes
- Cell Immortalization Step by Step
- Adherent and Suspension Cell Culture
- Cell Viability, Proliferation and Cytotoxicity Assays
- What Are CAR T Cells?
- From Blur to Clarity: Solving Resolution Limits in Live Cell Imaging
- Immunogenicity Testing: ELISA and MSD Assays
- Cultivated Meat: What to Know?
- Eosinophils vs. Basophils vs. Neutrophils
- Key Techniques in Primary, Immortalized and Stable Cell Line Development
- From Primary to Immortalized: Navigating Key Cell Lines in Biomedical Research
- Optimization Strategies of Cell-Based Assays
- 3D-Cell Model in Cell-Based Assay
- What Are the Pros and Cons of Adoptive Cell Therapy?
- 3D-Cell Model in Cell-Based Assay
- How to Maximize Efficiency in Cell-Based High-Throughput Screening?
- What are White Blood Cells?
- Immunogenicity Testing: ELISA and MSD Assays
- Types of Cell Therapy for Cancer
- Role of Cell-Based Assays in Drug Discovery and Development
- A Complete Guide to Immortalized Cancer Cell Lines in Cancer Research
- Exploring Cell Dynamics: Migration, Invasion, Adhesion, Angiogenesis, and EMT Assays
- Mastering Cell Culture and Cryopreservation: Key Strategies for Optimal Cell Viability and Stability
- Understanding Immunogenicity Assays: A Comprehensive Guide
-
Histology
- Fluorescent Nuclear Staining Dyes
- Stains Used in Histology
- Troubleshooting in Fluorescent Staining
- Immunohistochemistry Controls
- Overview of the FFPE Cell Pellet Product Lines
- Tips for Choosing the Right Protease Inhibitor
- Comparison of Membrane Stains vs. Cell Surface Stains
- Microscope Platforms
- How to Apply NGS Technologies to FFPE Tissues?
- Overview of Common Tracking Labels for MSCs
- Multiple Animal Tissue Arrays
- Cell and Tissue Fixation
- Immunohistochemistry Troubleshooting
- Cell Lysates: Composition, Properties, and Preparation
- Guides for Live Cell Imaging Dyes
- Instructions for Tumour Tissue Collection, Storage and Dissociation
- Mitochondrial Staining
- How to Choose the Right Antibody for Immunohistochemistry (IHC)
- How to Begin with Multiplex Immunohistochemistry (mIHC)
- Common Immunohistochemistry Stains and Their Role in Cancer Diagnosis
- Histological Staining Techniques: From Traditional Chemical Staining to Immunohistochemistry
- How Immunohistochemistry Makes the Invisible Brain Visible?
- Multiplexing Immunohistochemistry
- Modern Histological Techniques
- What You Must Know About Neuroscience IHC?
- Comparing IHC, ICC, and IF: Which One Fits Your Research?
- From Specimen to Slide: Core Methods in Histological Practice
- Serum vs. Plasma
-
Exosome
- Emerging Technologies and Methodologies for Exosome Research
- How to characterize exosomes?
- Classification, Isolation Techniques and Characterization of Exosomes
- Common Techniques for Exosome Nucleic Acid Extraction
- How Important are Lipids in Exosome Composition and Biogenesis?
- Collection of Exosome Samples and Precautions
- Current Research Status of Milk Exosomes
- How do PELN Deliver Drugs?
- What are the Functions of Exosomal Proteins?
- Exosome Size Measurement
- Production of Exosomes: Human Cell Lines and Cultivation Modes
- Techniques for Exosome Quantification
- Exosomes as Emerging Biomarker Tools for Diseases
- How to Apply Exosomes in Clinical?
- How to Perform Targeted Modification of Exosomes?
- Exosome Transfection for Altering Biomolecular Delivery
- Summary of Approaches for Loading Cargo into Exosomes
- Exosome Antibodies
- Exosome Quality Control: How to Do It?
- How to Enhancement Exosome Production?
- Applications of MSC-EVs in Immune Regulation and Regeneration
- The Role of Exosomes in Cancer
- What's the Potential of PELN in Disease Treatment?
- How to Efficiently Utilize MSC Exosomes for Disease Treatment?
- How to Label Exosomes?
- Unraveling Biogenesis and Composition of Exosomes
-
ISH/FISH
- ISH probe labeling method
- Multiple Approaches to Karyotyping
- RNAscope ISH Technology
- What are the Differences between FISH, aCGH, and NGS?
- CARD-FISH: Illuminating Microbial Diversity
- Comprehensive Comparison of IHC, CISH, and FISH Techniques
- In Situ Hybridization Probes
- Overview of Common FISH Techniques
- Guidelines for the Design of FISH Probes
- Small RNA Detection by ISH Methods
- Differences Between DNA and RNA Probes
- Overview of Oligo-FISH Technology
- Comparative Genomic Hybridization and Its Applications
- How to Use FISH in Hematologic Neoplasms?
- What Is the Use of FISH in Solid Tumors?
- FISH Tips and Troubleshooting
- Multiple Options for Proving Monoclonality
- FISH Techniques for Biofilm Detection
- Whole Chromosome Painting Probes for FISH
- Telomere Length Measurement Methods
- What Types of Multicolor FISH Probe Sets Are Available?
- Different Types of FISH Probes for Oncology Research
- Reagents Used in FISH Experiments
- Mapping of Transgenes by FISH
- What are Single, Dual, and Multiplex ISH?
- ImmunoFISH: Integrates FISH and IL for Dual Detection
- 9 ISH Tips You Can't Ignore
-
Toxicokinetics & Pharmacokinetics
- Toxicokinetics vs. Pharmacokinetics
- How to Improve Drug Plasma Stability?
- How Is the Cytotoxicity of Drugs Determined?
- How to Improve the Pharmacokinetic Properties of Peptides?
- Key Considerations in Toxicokinetic
- Organoids in Drug Discovery: Revolutionizing Therapeutic Research
- What Are Metabolism-Mediated Drug-Drug Interactions?
- Key Factors Influencing Brain Distribution of Drugs
- The Rise of In Vitro Testing in Drug Development
- Effects of Cytochrome P450 Metabolism on Drug Interactions
- Predictive Modeling of Metabolic Drug Toxicity
- Overview of In Vitro Permeability Assays
- How to Improve Drug Distribution in the Brain
- Organ-on-a-Chip Systems for Drug Screening
- Experimental Methods for Identifying Drug-Drug Interactions
- Traditional vs. Novel Drug Delivery Methods
- What factors influence drug distribution?
- How to Design and Synthesize Antibody Drug Conjugates?
- What Is the Role of the Blood-Brain Barrier in Drug Delivery?
- Parameters of Pharmacokinetics: Absorption, Distribution, Metabolism, Excretion
- Physical and Chemical Properties of Drugs and Calculations
- How to Conduct a Bioavailability Assessment?
- Comparison of MDCK-MDR1 and Caco-2 Cell-Based Permeability Assays
- Unraveling the Role of hERG Channels in Drug Safety
- Methods of Parallel Artificial Membrane Permeability Assays
- Pharmacokinetics Considerations for Antibody Drug Conjugates
- What are the Pharmacokinetic Properties of the Antisense Oligonucleotides?
- What Are Compartment Models in Pharmacokinetics?
- Pharmacokinetics of Therapeutic Peptides
- The Essentials of Quantitative Bioanalysis
- The 8 Costliest Mistakes in Preclinical CYP Phenotyping
- How Are Biomarkers Validated in Drug Development?
- Biomarkers vs. Functional Assays: Closing the Preclinical Gap
- When Should You Introduce ADME Tox Testing in Drug Development?
- 6 Easy Steps to Get Your In Vitro ADME Done
- From Cells to Systems: Modern Approaches to Disease Modeling
- How Can You Optimize Drug Toxicity Assessment?
- How to Choose the Right In Vitro ADME Assays for Small-Molecule Drugs
- What Are Biomarkers in Drug Discovery?
- The Bioanalysis Masterclass: Labile Metabolites
- How Genotoxicity Testing Guides Safer Drug Development
- Top 5 Pitfalls in In Vitro ADME Assays and How to Avoid Them
- 2D vs 3D Cell Culture Models: Which Is Best for Drug Toxicity Testing?
- Bioanalysis Errors: How to Spot and Fix Them Early
- Troubleshooting Common Issues in Drug Toxicity Testing
- What Is Genotoxicity in Pharmacology? Mechanisms and Sources
- Preclinical Workflow for Drug Toxicity Testing
- In Vitro ADME vs In Vivo ADME
- A Complete Guide to CYP Reaction Phenotyping in 2026
- How to Interpret CYP Phenotyping Data
- Reaction Phenotyping vs. Metabolic Stability
- The 8 Types of Drug Toxicity Every Researcher Must Know
- Why Cardiotoxicity Matters in R&D?
- What Are the Best Methods to Test Cardiotoxicity?
-
Disease Models
- Animal Models of Neurodegenerative Diseases
- Disease Models of Diabetes Mellitus
- Summary of Advantages and Limitations of Different Oncology Animal Models
- What Human Disease Models Are Available for Drug Development?
- Overview of Cardiovascular Disease Models in Drug Discovery
- Why Use PDX Models for Cancer Research?
- Preclinical Models of Acute Liver Failure
- Why Oncology Organoids Fail? How to Build Models That Work
- Organ-on-a-Chip: Is Your Microfluidic Setup Ready for Preclinical Trials?
- Oncology Model Strategy: From Screening to Validation
- How to Select the Right Preclinical Model for Drug Development
-
Cell Biology
- Life Science Articles
- Download Center
- Trending Newsletter