TUNEL Apoptosis Assay (Chromogenic)
Introduction
DNA fragmentation is a hallmark of late stage apoptosis, which can be detected in fixed cells and tissue sections by the terminal deoxynucleotidyl transferase (TdT) mediated dUTP nick-end labeling (TUNEL) assay. The TUNEL assay relies on the TdT enzyme to catalyze the addition of labeled dUTP to the 3’-hydroxyl (3’-OH) ends of cleaved DNA fragmentations. Labeled dUTP such as biotin-dUTP can be recognized using secondary reagent streptavidin through colorimetric detection. In addition, fluorescein-conjugated dUTP can be used for direct detection of fragmented DNA by flow cytometry or fluorescence microscopy.
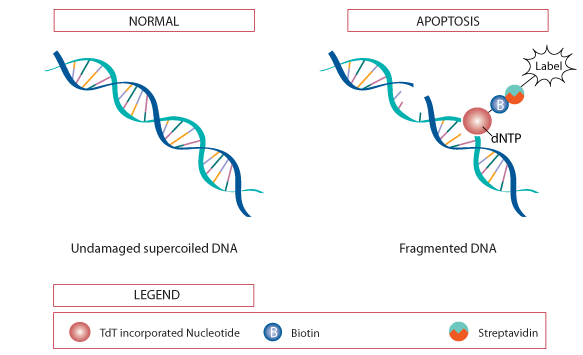 Figure 1. The principle of TUNEL assay.
Figure 1. The principle of TUNEL assay.
Preparation of Cultured Cells and Frozen Tissue Sections
- Adherent cells are collected and transferred to 6-well plates at a concentration of 105 cells/mL. Add apoptosis inducer to the cells and incubate according to your protocol.
- Suspension cells are harvested and centrifuged, after which the supernatant is discarded and the cells are resuspended to a concentration of approximately 2×107 cells/mL. Place 50-100 μL of the cell suspension on a polylysine-coated slide and then move around to form a thin and even film with a glass spreader.
- Conventional method is used for making fresh-frozen tissue sections.
- Wash cells or sections twice with PBS.
- Add sufficient fixative (4% paraformaldehyde) to complete cover the cells and sections (not required for fixed-frozen sections).
- Incubate for 30 minutes at 4ºC.
- Remove fixative and wash twice in PBS.
- Permeabilize in 0.2% Triton X-100 for 30 minutes at room temperature.
- Wash twice with PBS.
Note: Apoptotic cells maybe detach from adherent cell cultures and lost during wash steps, we recommend fixing directly without performing the wash steps.
Preparation of Paraffin Tissue Sections
- The paraffin tissue sections are deparaffinized at room temperature according to Tabel 1.
Table 1. Tissue deparaffinization procedure.
Xylenes Xylenes 100% EtOH 100% EtOH 95% EtOH 85% EtOH 75% EtOH 50% EtOH 1X PBS 5 min 5 min 5 min 5 min 5 min 3 min 3 min 3 min 5 min - Permeabilize paraffin sections with 100 µL proteinase K solution (20 µg/mL) for 30 minutes at room temperature.
- Incubation time and temperature may require optimization depending on tissue types.
- Rinse in PBS.
Positive Control Preparation
The DNase I generates strand breaks in DNA which provide a positive TUNEL reaction.
- Wash sample with deionized water.
- Prepare DNase I solution according to Table 2 and mix well.
Table 2. DNase I solution.
Reaction components Number of Samples 1 2 3 Deionized water 89 µL 178 µL 267 µL DNase I buffer 10 µL 20 µL 30 µL DNase I 1 µL 2 µL 3 µL Total volume 100 µL 200 µL 300 µL - Incubate samples for 20 minutes at room temperature with 100 µL DNase I solution.
- Wash once with deionized water and then proceed to TdT reaction.
Note: Do not vortex the DNase I solution due to DNase I denatures with vigorous mixing.
TUNEL Reaction
- Incubate samples with 2% hydrogen peroxide for 5 min at room temperature to inactivate endogenous peroxidases.
- Wash 2 x 5 minutes with PBS.
- Incubate samples with 100 µL TdT reaction buffer for 10 minutes at room temperature.
- TdT reaction cocktails are prepared according to Table 3 before use.
Table 3. TdT reaction cocktails.
Reaction components Number of Samples 1 2 4 5 10 TdT reaction buffer 94 µL 188 µL 376 µL 470 µL 940 µL TdT enzyme 4 µL 8 µL 16 µL 20 µL 40 µL Biotin-dUTP 2 µL 4 µL 8 µL 10 µL 20 µL Total volume 100 µL 200 µL 400 µL 500 µL 1 mL - Remove TdT reaction buffer and add 100 µL TdT reaction cocktails to each sample, allowing the solution to cover the surface completely.
- For cell staining, incubate for 60 minutes at 37ºC. Tissue staining may require 2 hours.
- Stop the reaction by incubating samples in 2 X SSC.
- Wash samples 2 x 10 minutes in PBS with 3% BSA.
- HRP-streptavidin staining solution is prepared according to Table 4.
Table 4. HRP-streptavidin staining solution.
Reaction components Number of Samples 1 2 4 5 10 Staining buffer 99 µL 198 µL 396 µL 495 µL 990 µL HRP-streptavidin 1 µL 2 µL 4 µL 5 µL 10 µL Total volume 100 µL 200 µL 400 µL 500 µL 1 mL - Add 100 µL HRP-streptavidin staining solution to each sample, and incubate for 30 minutes at room temperature. Tissue staining may require 1 hour incubation.
- Wash samples 2 x 5 minutes in PBS with 3% BSA.
- DAB staining solution is prepared according to Table 5.
Table 5. DAB staining solution.
Reaction components Number of Samples 1 2 4 5 10 DAB diluent 97 µL 194 µL 388 µL 485 µL 970 µL DAB stock solution 3 µL 6 µL 12 µL 15 µL 30 µL Total volume 100 µL 200 µL 400 µL 500 µL 1 mL - Add 100 µL DAB staining solution to each sample, and incubate at room temperature.
- Monitor color development until desired level of staining is achieved. Stop the reaction by rinsing with PBS.
- Counterstain samples with hematoxylin stain if desired.
- Image and analyze in light microscopy.
Note: A humidified chamber is recommended to protect against evaporation.
Cell Services:
Cell Line Testing and Assays: