Immortalized Human Pancreatic Islet-Derived Precursor Cells-SV40
Cat.No.: CSC-I9029L
Species: Homo sapiens
Source: Pancreas
Morphology: Polygonal
Culture Properties: Adherent
- Specification
- Background
- Scientific Data
- Q & A
- Customer Review
Note: Never can cells be kept at -20 °C.
Immortalized Human Pancreatic Islet-Derived Precursor Cells - SV40 are a cell line that was generated from human pancreatic islet-associated precursor cells and immortalized with SV40 large T antigen allowing for extended proliferation in vitro. The cells represent a precocious, progenitor-like state of pancreatic cells that are not readily accessible for study through more mature islet cells.
Key characteristics of these cells include a precursor phenotype. These cells do not behave like terminally differentiated endocrine cells, but rather can be induced to proliferate and respond to other developmental cues and express known pancreatic progenitor and early endocrine commitment markers. This makes them an ideal system to study lineage specification, β-cell development, and molecular transitions between progenitor and mature islet cell states. These cells have been used extensively to study diabetes and pancreatic biology. They provide a stable and reproducible system to explore the mechanisms of islet regeneration, cellular reprogramming and β-cell replacement therapies. Their long term growth characteristics and human origin also make them an excellent system for screening to identify compounds that promote islet differentiation, survival or function bridging basic pancreatic biology and translational diabetes research.
Analysis of MicroRNA Signature Differentially Expressed in Pancreatic Islet Cells Treated with Pancreatic Cancer-Derived Exosomes
PC patients often develop insulin resistance or DM before diagnosis, highlighting PC-DM as a potential detection platform. Here, Kim et al. treated immortalized human pancreatic islet-derived precursor cells (hIPCs) with PC-derived exosomes (from 50 patients) and healthy-derived exosomes (from 50 individuals) and assessed miRNA expression changes using next-generation sequencing and bioinformatic analysis.
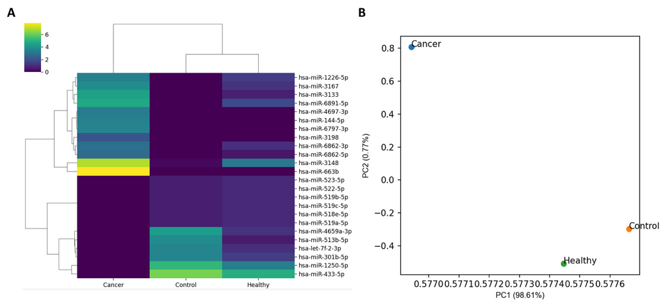
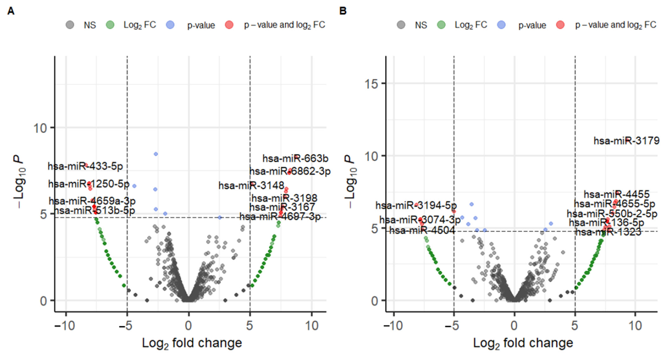
In the expression analysis, 2656 miRNA markers were used. The heatmap of normalized miRNA expression from three specimens and the scatter plot from principal component analysis (PCA) are shown in Figure 1. Unsupervised clustering analysis revealed that Control and Healthy clustered together, while Cancer was distant from both, indicating distinct expression profiles. The results of the differential expression analysis (DEmiRNA1 and DEmiRNA2) are shown in Figure 2. DEmiRNA1 (Cancer vs. Control) identified 13 upregulated and 16 downregulated miRNAs. DEmiRNA2 (Healthy vs. Control) identified 14 upregulated and 7 downregulated miRNAs. Among these, 1 miRNA was upregulated in both DEmiRNA1 and DEmiRNA2, and 4 miRNAs were downregulated in both. Thus, the candidate markers from the DEmiRNA analysis totaled 24, comprising 12 upregulated and 12 downregulated miRNAs.
Ask a Question
Write your own review
- Adipose Tissue-Derived Stem Cells
- Human Neurons
- Mouse Probe
- Whole Chromosome Painting Probes
- Hepatic Cells
- Renal Cells
- In Vitro ADME Kits
- Tissue Microarray
- Tissue Blocks
- Tissue Sections
- FFPE Cell Pellet
- Probe
- Centromere Probes
- Telomere Probes
- Satellite Enumeration Probes
- Subtelomere Specific Probes
- Bacterial Probes
- ISH/FISH Probes
- Exosome Isolation Kit
- Human Adult Stem Cells
- Mouse Stem Cells
- iPSCs
- Mouse Embryonic Stem Cells
- iPSC Differentiation Kits
- Mesenchymal Stem Cells
- Immortalized Human Cells
- Immortalized Murine Cells
- Cell Immortalization Kit
- Adipose Cells
- Cardiac Cells
- Dermal Cells
- Epidermal Cells
- Peripheral Blood Mononuclear Cells
- Umbilical Cord Cells
- Monkey Primary Cells
- Mouse Primary Cells
- Breast Tumor Cells
- Colorectal Tumor Cells
- Esophageal Tumor Cells
- Lung Tumor Cells
- Leukemia/Lymphoma/Myeloma Cells
- Ovarian Tumor Cells
- Pancreatic Tumor Cells
- Mouse Tumor Cells